In contemporary diagnostics and risk intelligence, it is essential to distinguish between technology and application. Next-Generation Sequencing (NGS) represents the technological platform, while Metagenomics is the applied scientific approach that leverages this technology to generate comprehensive environmental insight.
At METS Laboratory, metagenomics is best described as the “Identity Card” of an environment—a complete genetic blueprint that reveals which microorganisms are present, what biological functions they are capable of performing, and how they coexist and evolve within a given ecosystem.

Why the Industry Is Moving Beyond Traditional Microbiology
Conventional microbiological testing relies primarily on selective culturing techniques, which impose several critical limitations:
- Detection is restricted to microorganisms capable of growing under laboratory conditions
- Testing focuses only on predefined or expected pathogens
- Turnaround times are often extended due to incubation requirements
- More than 99% of environmental microorganisms, which are unculturable, remain undetected
As a result, traditional microbiology offers a narrow and incomplete snapshot of microbial presence.
Metagenomic sequencing, by contrast, captures the entire microbial DNA landscape in a single workflow. This enables faster analysis, deeper detection, and predictive insights, rather than delayed and reactive responses.
This transition reflects a broader industry shift—from compliance-driven testing to proactive microbial risk intelligence.

How Metagenomics Works at METS Laboratory
The metagenomic workflow at METS Laboratory integrates state-of-the-art sequencing platforms with advanced bioinformatics and scientific interpretation.
- Sample Collection & DNA Extraction
Environmental or product samples are carefully collected and processed to extract total microbial DNA, ensuring no bias toward any specific organism or microbial group.
- Next-Generation Sequencing (NGS)
Millions of DNA fragments are sequenced simultaneously using high-throughput NGS platforms, producing high-resolution genomic datasets representative of the complete microbial community.
- Bioinformatics & Data Interpretation
Proprietary bioinformatics pipelines analyze the sequencing data by comparing genetic signatures against curated and validated databases. This enables precise insight into:
- Microbial taxonomy
- Functional genes
- Relative abundance and microbial load
- Action-Oriented Reporting
Rather than delivering raw sequencing output alone, METS Laboratory translates complex genomic data into clear, actionable risk intelligence. Reports are designed to support informed decision-making across food safety, environmental monitoring, and quality assurance systems.

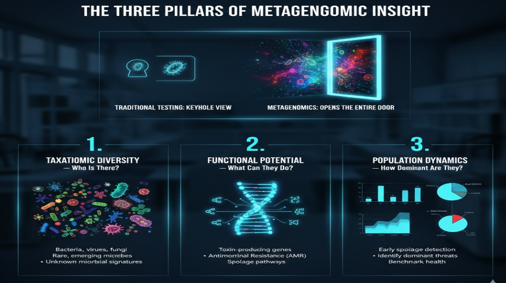
The Three Pillars of Metagenomic Insight
Where traditional microbiology provides a limited viewpoint, metagenomics offers complete visibility into microbial ecosystems.
- Taxonomic Diversity — Who Is There?
Metagenomics enables identification of the entire microbial community, including:
- Bacteria, viruses, fungi, and parasites
- Rare, emerging, and unculturable organisms
- Previously uncharacterized or unknown microbial signatures
This level of comprehensive detection is unachievable through culture-based methods alone.
- Functional Potential — What Can They Do?
Beyond identifying organisms, metagenomics reveals genetic functionality. By analyzing gene content, METS Laboratory can detect:
- Virulence-associated genes
- Toxin-producing capabilities
- Antimicrobial resistance (AMR) markers
- Metabolic pathways linked to spoilage, degradation, or biodegradation
This functional insight transforms microbial testing into a true risk assessment tool, rather than a simple presence/absence test.
- Population Dynamics — How Dominant Are They?
By quantifying relative abundance and microbial load, metagenomics allows for:
- Early detection of spoilage organism overgrowth
- Identification of dominant high-risk microorganisms before system failures occur
- Benchmarking and comparison of healthy versus unhealthy microbial ecosystems
Real-World Applications at METS Laboratory
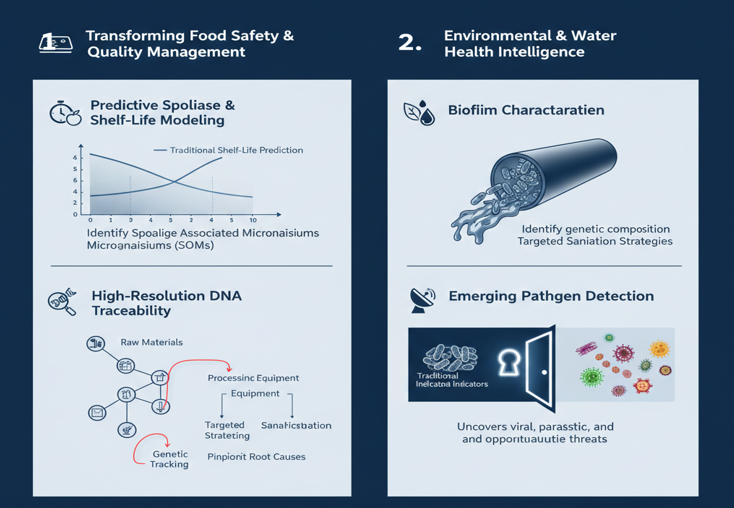
- Transforming Food Safety & Quality Management
Traditional food microbiology is reactive and target-specific—organisms not included in the test panel remain invisible. Metagenomics replaces this limitation with total microbial surveillance.
Predictive Spoilage & Shelf-Life Modeling (h4)
Identification of Spoilage Associated Microorganisms (SOMs) enables manufacturers to predict product deterioration before sensory defects become apparent. This proactive insight reduces recalls, minimizes food waste, and protects brand integrity.
High-Resolution DNA Traceability (h4)
In contamination investigations, metagenomics enables genetic source tracking. By matching microbial fingerprints across raw materials, processing equipment, and finished products, METS Laboratory can precisely identify contamination origins and transmission pathways.
- Environmental & Water Health Intelligence
Metagenomics functions as an advanced biosurveillance system for complex environmental matrices where traditional indicator organisms are insufficient.
Biofilm Characterization
Biofilms are highly structured microbial communities with enhanced resistance to cleaning agents and disinfectants. Metagenomic analysis reveals their genetic composition, enabling the development of targeted and effective sanitation strategies rather than trial-and-error approaches.
Emerging Pathogen Detection
In water and environmental monitoring programs, metagenomics detects viral, parasitic, and opportunistic microbial threats that are often missed when relying solely on standard indicators such as coliforms.
Why This Matters for the Industry
The future of food safety, environmental monitoring, and regulatory compliance is genomics-driven. By transitioning from a “grow-and-count” paradigm to a “sequence-and-analyze” approach, organizations achieve:
- Faster and more comprehensive insights
- Broader and unbiased detection capabilities
- Predictive and preventive risk management
- Enhanced brand protection and operational confidence
METS Laboratory is at the forefront of this transformation—delivering scientifically robust, data-rich metagenomic solutions that enable industries to move from reaction to prevention, and from compliance to confidence.
